Predicting the 3D structure of proteins remains one of the hardest problems in computational biology — NP‑complete even in simplified models. While deep learning (AlphaFold) and citizen science (FoldIT) have made tremendous progress, many cases (intrinsically disordered proteins, membrane proteins, metalloenzymes) still require quantum‑aware energy evaluations and human intuition.
qFoldIT is an open platform that unifies three complementary approaches: DeepFoldIT (neural network initializer), Rosetta@Home (classical Metropolis), and qFold (quantum‑accelerated Metropolis). Gamification invites thousands of players to contribute, while AAA‑grade visualization (Unigine 2 Sim) makes molecular interactions tangible.
The platform is built on Nobel‑prize winning discoveries and is entirely free for academic use.
qFoldIT integrates six Nobel‑awarded breakthroughs, grouped by scientific theme:
Physics 2024 (Hopfield, Hinton) — neural networks and associative memory. Used in qFoldIT for adaptive AI assistants, procedural generation of Arctic landscapes, and molecular structure prediction.
Chemistry 2024 (Baker, Hassabis, Jumper) — de novo protein design and AlphaFold. Our DeepFoldIT module and quantum‑accelerated folding build directly on these advances.
Chemistry 2013 (Karplus, Levitt, Warshel) — multiscale models for complex chemical systems, underpinning our hybrid quantum‑classical dynamics.
Physics 2025 (Clarke, Devoret, Martinis) — large‑scale quantum systems. The quantum‑adapter module provides cloud‑based quantum annealing and VQE calculations for molecular folding, directly leveraging this legacy.
Chemistry 2025 (Kitagawa, Robson, Yaghi) — metal‑organic frameworks. qFoldIT extends its molecular design engine to porous materials for carbon capture, catalysis, and hydrogen storage.
Medicine 2000 (Kandel) — long‑term potentiation. Our core game loop uses immediate feedback and spaced repetition to strengthen neural pathways.
Chemistry 2018 (Arnold) — directed evolution of enzymes. Players mutate and select protein variants in a gamified evolution loop, evaluated by ZairaChem AI.
Peace 2007 (IPCC, Gore) — climate change awareness. qFoldIT is the digital twin of the Arctic station “Snezhinka” (AHEAD project, Arctic Council), where players design carbon‑free technologies and bioremediation enzymes.
Economics 2002 (Kahneman) — two systems of thinking (intuitive System 1 vs analytical System 2). qFoldIT dynamically switches between high‑action Arctic races (System 1) and deep molecular folding puzzles (System 2), preventing fatigue and maintaining flow.
Five world‑class game designers and visual artists shape the qFoldIT experience:
- George Ahiakwa (founder of qFoldIT) — developer on Atomic Heart (Mundfish).
- Demis Hassabis — lead programmer on Theme Park (Bullfrog), lead AI on Black & White (Lionhead), founder of Elixir Studios (Republic: The Revolution, Evil Genius).
- Dr. Beata Mierzwa — creator of the award‑winning educational game Microscopya.
- Robertson Nortey — Lead Gameplay & Systems Designer (Rocksteady Studios: Batman Arkham series, Suicide Squad; currently at Leti Arts). Brings AAA combat, AI, and open‑world expertise to make molecular folding feel like a blockbuster action puzzle.
- Karen Happuch — founder of KHPH Studios (Ghana), director and animator of the acclaimed short Abrefi Kɔtɔ. She is the visual storyteller who translates Adinkra symbols and African fractal logic into the interactive visual language of qFoldIT, ensuring the game's cultural soul shines at every scale — from 2D pixel art to cinematic VR.
From Atomic Heart to qFoldIT: Working on Atomic Heart was built around “science fiction” — an opportunity to creatively fantasise about the visual realisation and physical behaviour of substances (e.g., the “polymer”). That experience taught how to translate abstract physical properties into compelling, visceral gameplay. qFoldIT applies the same philosophy: molecular interactions, hydrogen bonds, and quantum tunnelling become visible, tangible, and playable — turning hard science into an intuitive experience.
Dr. Beata Mierzwa’s game Microscopya is a scientifically accurate adventure inside a living cell, hand‑drawn from real microscopy data. It has received multiple prestigious awards:
- James Paul GEE! Learning Game Awards (2022) — winner in “Amateur Created” and “People’s Choice”.
- International Educational Games Competition (ECGBL 2022) — second place.
- Serious Play Conference Awards — recognised for excellence.
- Media coverage: The Scientist Magazine, Science Magazine, Nerdist, KPBS News.



This recognition from Microsoft validates LetiArts’ leading role in African game innovation. The same creative and technical excellence powers qFoldIT’s gamified molecular design, blending AAA production values with cutting-edge science. Inspired by world‑class scientific animators like Drew Berry (whose breathtaking visualizations of molecular machines and quantum biology set the gold standard), qFoldIT brings the same cinematic wonder to interactive protein folding. The Microscopya poster above captures the artistic rigor and biological accuracy that define our approach — turning the invisible into an unforgettable experience.
The same design principles — artistic rigour, biological accuracy, and engaging interactivity — drive qFoldIT. Players not only play but actively contribute to molecular design, with their solutions validated by AI (ZairaChem) and quantum algorithms.
qFoldIT integrates three complementary computational engines:
- DeepFoldIT — a deep residual network (inspired by AlphaFold) predicts torsion angles (φ, ψ) and distances, generating initial conformations.
- Rosetta@Home — classical Metropolis Monte Carlo with the Rosetta energy function performs local optimisation on CPU clusters.
- qFold — a coined Szegedy quantum walk (quantum Metropolis) that provides quadratic speedup for rugged energy landscapes. It runs on simulators (Qiskit‑Aer) or real quantum backends (IonQ, D‑Wave, Rosatom).
qFoldIT solves the fundamental trade‑off of molecular editors: beautiful but imprecise (Unity/Unreal) vs. accurate but cumbersome (command‑line tools). Our stack combines UNIGINE's 64‑bit precision, NASA‑derived interaction logic, real‑time synchronization via NanoVer, and cloud quantum acceleration into a seamless, gamified experience.
NanoVer is more than an iMD engine — it is the real‑time translation layer that makes the 4‑layer stack possible. It decouples the fast visual feedback (UNIGINE) from the slower, more accurate quantum evaluations (Qiskit). Every atomic displacement is streamed via gRPC, logged with scientific precision, and immediately reflected in collaborative sessions.
- Live force feedback — Players can “grab” atoms, apply forces in real time, and see the molecular response within milliseconds.
- Quantitatively accurate iMD — Every perturbation is recorded, making exploration fully reproducible.
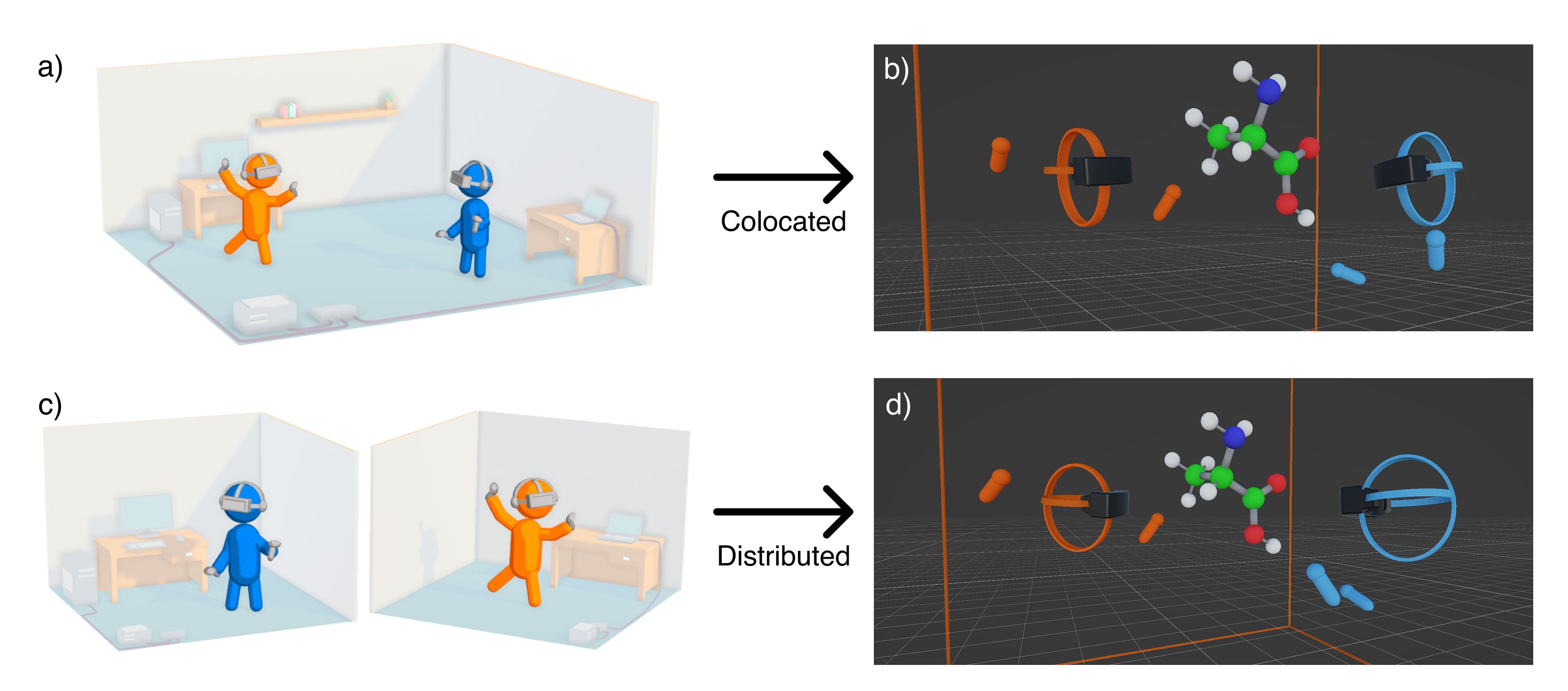
- Multi‑user colocated/distributed VR — Share the same virtual environment and collaboratively fold proteins.
- Jupyter‑native analysis — Real‑time plotting of energies bridges gamification with data science.
From city‑farms and microscopes to space‑grade quantum hardware — qFoldIT extends its digital‑twin philosophy to the Cold Atom Lab (CAL) aboard the ISS. CAL creates Bose‑Einstein condensates, “giant atoms” that make quantum effects visible on a macroscopic scale.

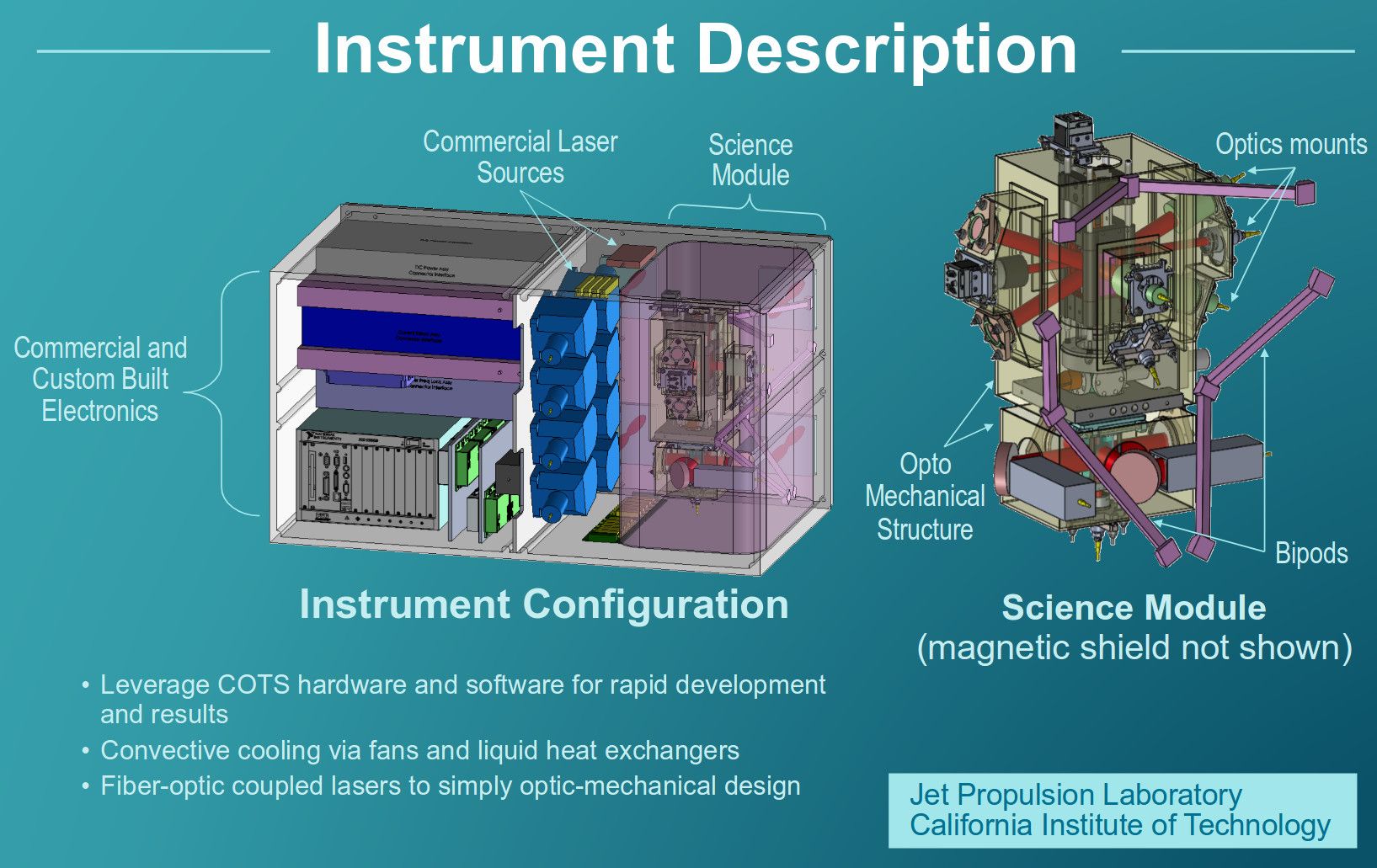

In qFoldIT, the CAL module serves as a high‑fidelity training simulator — a direct evolution of the platform’s proven ability to control real‑world equipment. Players align laser beams, tune magnetic traps, and replace modules using holographic guides.
All modules — DeepFoldIT, Rosetta, quantum‑adapter, NanoVer Server — run as Docker containers orchestrated by Kubernetes on Amazon EKS. This ensures GPU‑aware scaling, high availability, and strict separation of classical, quantum, and interactive workloads. All code is planned for open release on GitHub.
Additional partners: Joint Institute for Nuclear Research (JINR, Dubna) and Yonsei University (Seoul, Korea) have joined as administrative collaborators, contributing expertise in machine learning, computational chemistry, and generative AI for protein‑protein interactions.
We invite: computational biologists, quantum algorithm researchers, game developers, and citizen scientists. Beta access is free — contact us to participate in the closed beta (planned for late 2026).

qFoldIT embodies the digital‑to‑biological convergence: DNA → computer → “living factory”. This enables drugs tailored to genetic diversity, DNA‑based data storage, and distributed biomanufacturing — all within an open, gamified framework.







